GSoC 2021 : SBML4Humans - Interactive SBML Report for Humans - Final Report
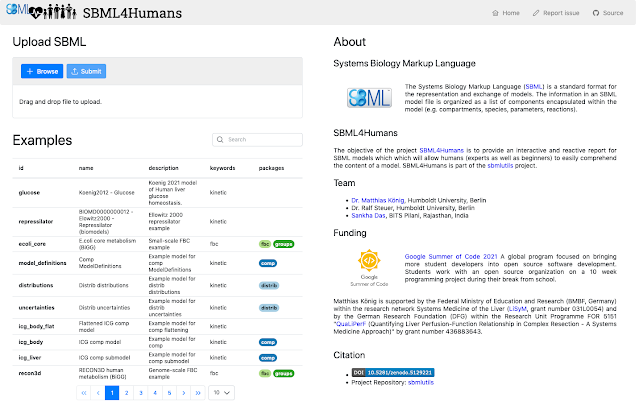
Overview Over the past 10 weeks, me and my mentors, Dr. Matthias König and Dr. Ralf Steuer have been working on the project “SBML4Humans - Interactive SBML Report for Humans” under Google Summer of Code 2021. The project is headed by the organization National Resource for Network Biology (NRNB). As the programme comes to an end this week, the project is also nearing its successful completion. The final product created out of this project is an interactive web application that generates human-readable, customizable reports for biological models written in SBML, now available at https://sbml4humans.de . The web application was developed using the Vue.js framework and uses the sbmlutils package written in Python as a backend. An API instance created using the FastAPI package provides endpoints to make requests to the sbmlutil s package from the SBML4Humans application. The basic workflow of the application is as follows: An SBML file is uploaded to the client web application and...